Similarity search, a key concept in data science, enables researchers to analyse huge volumes of unstructured data that is not accessible by conventional query search engines.
The technique entails examining the bit-level differences between millions or billions of database records by finding content that is similar to each other. Use cases include face recognition, DNA sequencing, researching drug candidate molecules, SHA1 algorithm hashes and natural language processing (NLP). Facebook’s FAISS library is a prominent example of similarity search in action.
Typically, Xeon CPUs and GPUs are pressed into service to process similarity searches but neither technology is optimised for this work. When handling very large datasets there is a memory-to-CPU-core bottleneck.
A Xeon CPU can search one record at a time per core. When a Xeon CPU runs a similarity search, looking for occurrences of a search term in a target dataset, the dataset or portions of it are read into memory and the Xeon core, or cores, compares each dataset entry with the search term. If the dataset is an image recognition database containing one billion records that search can take a long time. Also, this has an implication for power requirements.
A Nvidia GPU can throw more cores at the problem but even that takes too long when a billion-record database is involved.
So says GSI Technology, a Silicon Valley speciality memory maker, which has designed a parallel processing system dedicated to the single similarity search job. The company claims its Gemini ‘associative processing unit’ (APU) conducts similarity searches on ‘certain big data workloads’ 100 times faster than standard Xeon processors, while reducing power by 70 per cent.
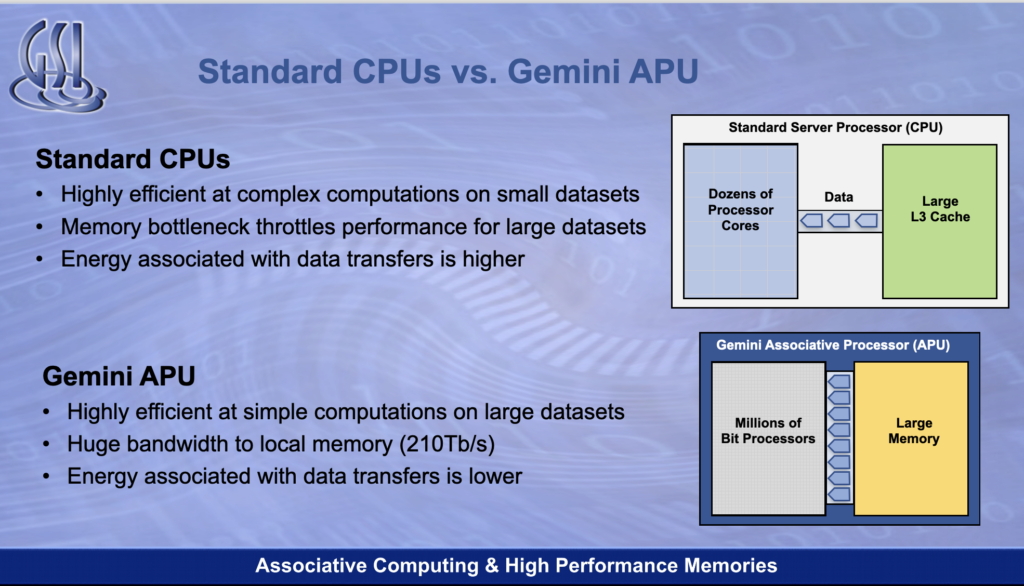

The APU locates compute units directly in a memory array so they can process data in parallel. This avoids moving the data from memory to a server’s Xeon CPU core, traversing Level 3, 2, and 1 caches on the way.
GSI’s Gemini APU accelerator is deployed as a server offload card. Consider this as an example of a data processing unit (DPU). It performs a specific function much faster than a server’s X86 processors, and can be used to offload it, freeing it to run applications in virtual machines and containers.
According to the company, a GSI card with four onboard Gemini APUs found the matches for a scanned face in a billion-record database in 1.25 milliseconds. The records were quite large, with each containing a set of 768-bit vectors hashed from 96 facial image features. A Xeon server takes up to 125 milliseconds to do the same search, according to GSI.
The company said a 1U server with 16 Gemini chips performed 5.4 million hashes per second when running the 256-bit SHA1 algorithm. This is greater throughput than a 4U server holding eight Nvidia V100 cards and uses half the electrical power, according to GSI.
We have no price for a Gemini APU but single quantity pricing for GSI’s Leda-branded PCIe card with four Geminis mounted on it is $15,000.
GSI’s APU
The Gemini APU combines SRAM and two million bit-processors for in-memory computing functions. SRAM – short for Static Random Access Memory – is faster and more expensive than DRAM.
The GSI interweaves 1-bit processing units with the read-modify-write lines of SRAM in its Gemini chip. All these processors work in parallel.
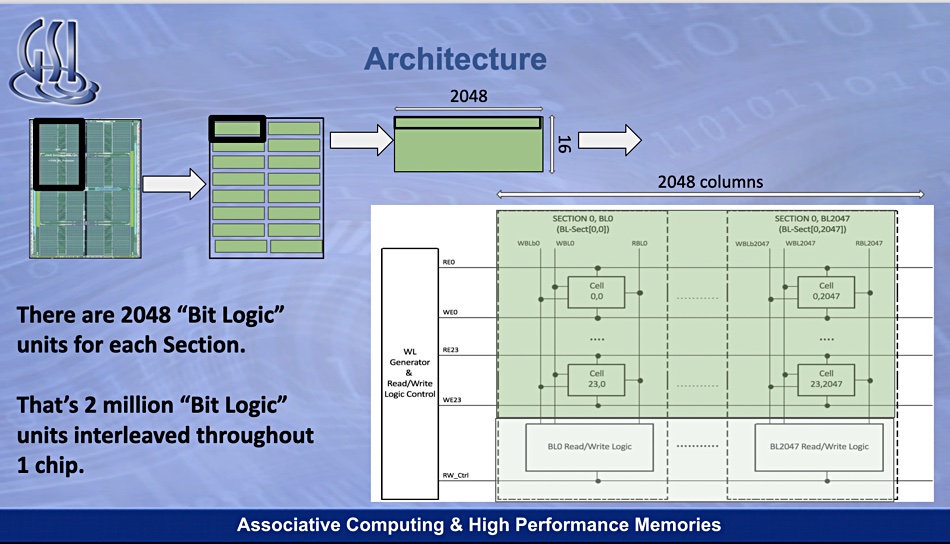
In a Gemini chip, data flows directly from the memory cells into a nearby chip and the search term is loaded onto each processor. This is compared to the string loaded from the SRAM cells, with the two million-plus cores operating in parallel to compute Hamming* distance, greatly outperforming a Xeon core, or even 28 Xeon cores, doing the same work.
A Gemini chip can perform two million x 1-bit operations per 400MHz clock cycle with a 26 TB/sec memory bandwidth, whereas a Xeon 8280 can do 28 x 2 x 512 bits at 2.7GHz with a 1TB/sec memory bandwidth.
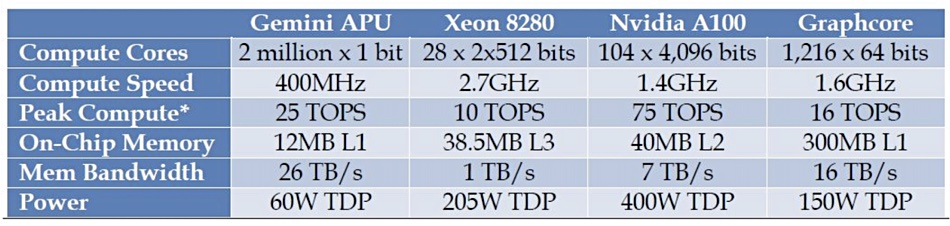
For comparison, a Nvidia A100 GPU server can complete 104 x 4,096 bits per 1.4GHz clock cycle, providing a 7TB/sec memory bandwidth. The Gemini chip’s memory bandwidth leaves the A100 trailing in the dust, and the Xeon CPU is even further behind.
*Hamming Distance When a computer runs a search it deals with a search term, represented as a binary string, and it looks for equivalent or similar search terms. The similarity can be expressed as the difference between the search term string and a target string, described as the number of bit positions in which the two bits are different.
It works like this; envisage two strings of equal length; 1101 1001 and 1001 1101. Add them together, 11011001 + 10011101, to get = 01000100. This contains two 1s and the Hamming distance is 2. Other things being equal, strings with smaller Hamming distances are more likely to represent things that are similar than strings with greater Hamming distances. The ‘things’ can be facial recognition images, genomes, drug candidate molecules, SHA1 algorithm hashes and so forth.