A researcher at the Rochester Institute of Technology (RIT) has devised a microfluidic lab on a chip that can perform artificial neural network (ANN) computation on data stored in DNA.
DNA storage relies on data being stored as specific combinations of the four nucleobases – cytosine (C), guanine (G), adenine (A), and thymine (T) – found in the double helix formation in the DNA biopolymer molecule. One method has pairs of these nucleobases symbolizing binary ones and zeros. Amlan Ganguly, computer engineering department head in RIT’s Kate Gleason College of Engineering, co-authored a scientific paper in which he envisions “a computing platform using DNA molecules that are capable of computation in-situ without the need for domain conversion of information from DNA to electronics.”
Ganguly states: “DNA is excellent at storing information, in fact, it is much better than the electronic modes of memory because it is about 3-to-6 orders of magnitude more compact than most memory hardware that we have; it is also much more reliable and durable.”
He adds: “We proposed to represent numbers through concentrations of solutions containing specifically manipulated DNA molecules and computing operations as manipulation of DNA molecules – operations like addition and multiplication and other non-linear functions necessary for network computations can be performed. That is the bridge from storage to computation and using DNA as a vehicle to do the computation.”
The paper states: “While biochemical reaction representing computations using DNA molecules are several orders of magnitude slower than electronic gates, their data density is 3 orders of magnitude higher and 8 orders of magnitude lower in energy consumption than solid state memory.”
The RIT integrated circuit (IC) based on microfluidics is suited for “highly dense, throughput-demanding bio-compatible applications such as an intelligent Organ-on-Chip or other biomedical applications that may not be latency-critical.”
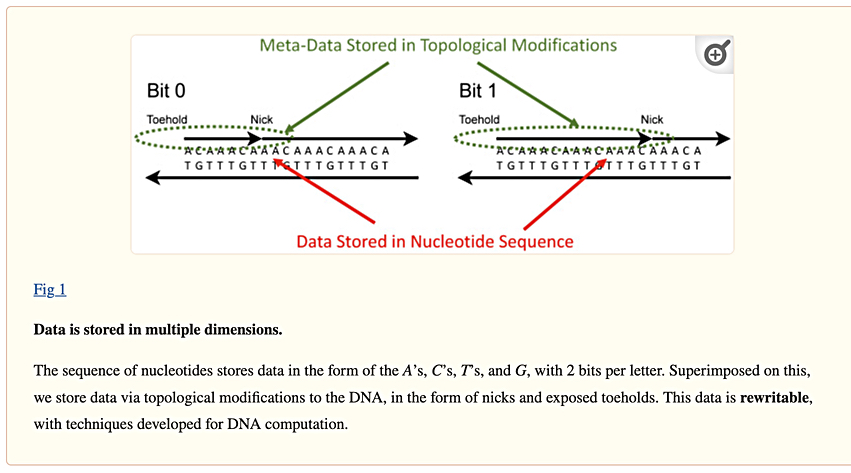
The research paper abstract says: ”It computes entirely in the molecular domain without converting data to electrical form, making it a form of in-memory computing on DNA. The computation is achieved by topologically modifying DNA strands through the use of enzymes called nickases.”
Data is represented stochastically through the concentration of the DNA molecules that are nicked at specific sites. A stochastic process has random or probabilistic dynamics over time, and the randomness is modeled mathematically to study the statistical properties of the process. The probabilities of different outcomes can be analyzed.
Ganguly says DNA computation and storage uses less energy than electronic storage and computation: “We are in the age of big data that needs to be stored somewhere. We don’t think that more datacenters are the answer, or even the best answer. Each datacenter requires the equivalent of a city block of power. Building, maintaining, and operating more traditional datacenters is not sustainable.”
Comment
DNA storage has little chance of replacing electronic storage media such as disk and solid state drives, which have millisecond-class data access speeds, and DRAM and CPU/GPU computation operate at microsecond-class speeds or faster. This is because reading and writing with DNA, meaning sequencing and synthesizing DNA molecules, is slow. It needs hours. Ganguly admits that computation using biochemical reactions is several orders of magnitude slower than using a CPU or GPU.