French startup Biomemory has developed a DNA Drive storage technology using synthetic biology to store 5PB of data in stainless steel pellets with contents that can be read by a DNA sequencer.
Biomemory briefed an IT Press Tour in Paris in September about its technology, which is intended to replace tape-based archives. The DNA Drive is basically the equivalent of a tape cartridge in that it has no on-board data writing or reading capability, such as a disk drive or SSD; it is a dumb data container containing up to 100 billion copies of a file. It is intended for use in automated library devices, equivalent to tape autoloaders or library systems with robots.

The company describes the age-old use case: data growing faster than the ability of the storage industry to store it. Worldwide data amounted to 45ZB in 2020, according to tech analyst IDC, which predicts this will grow to 175ZB in 2025. Some 70 percent of this will be archive data mostly stored on tape, which has a 15-25 year lifespan. DNA stored in airless and lightless conditions is said to be able to last for centuries, if not thousands of years.
DNA is a biopolymer molecule composed from two chains in a double helix formation, and carrying genetic information. The chains are made up from nucleotides containing one of four nucleobases; cytosine (C), guanine (G), adenine (A) and thymine (T). There is a density of 50 atoms per letter. Both chains carry the same data, which is encoded into sequences of the four nucleobases. There is 1 bit per base in Biomemory’s coding scheme, attributing randomly A or C for 0 and T or G for 1. A twist of DNA occupies a 2nm by 3.4nm physical space.
In our human genome there are 6.4 billion pairs of nucleotides per cell, equivalent to 700 MB/cell. A human being contains around 3,900 billion cells, which is equivalent to 2.7ZB of data. Biomemory claims all of the world’s digital data in 2020, 45ZB of it, could be stored in one gram of DNA.
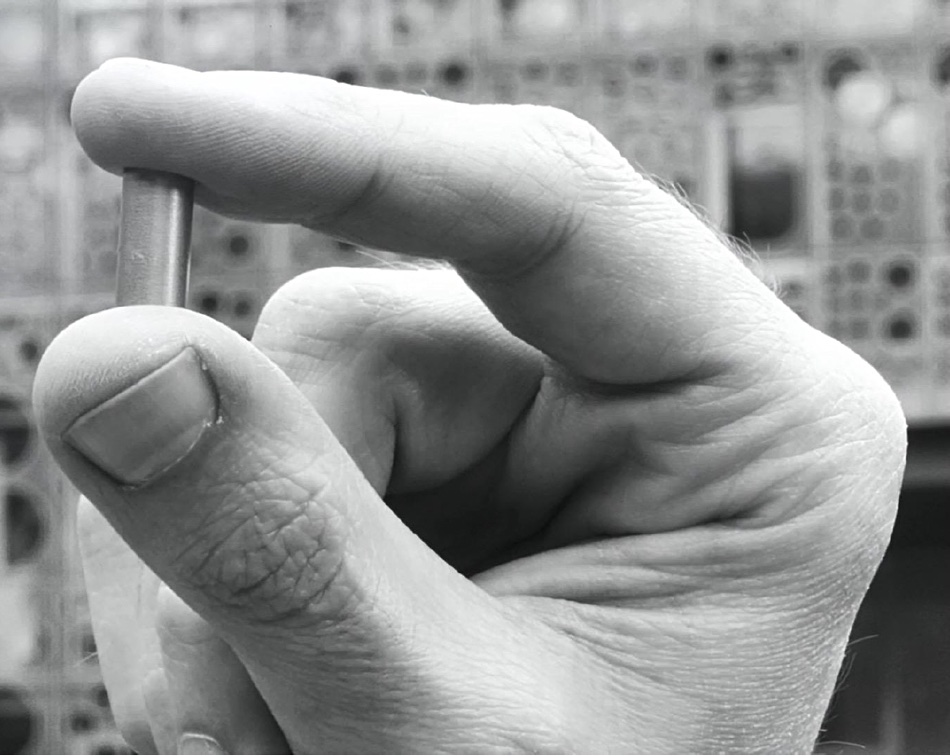
The data writing process involves coding and compressing the data into 4-brick DNA strings, creating the double-stranded DNA molecules, inserting them into Escherichia coli (E. coli) bacteria, getting them to reproduce themselves and hence replicate the DNA strands, extracting and then purifying and drying the DNA, inserting it into the capsules, and sealing them.
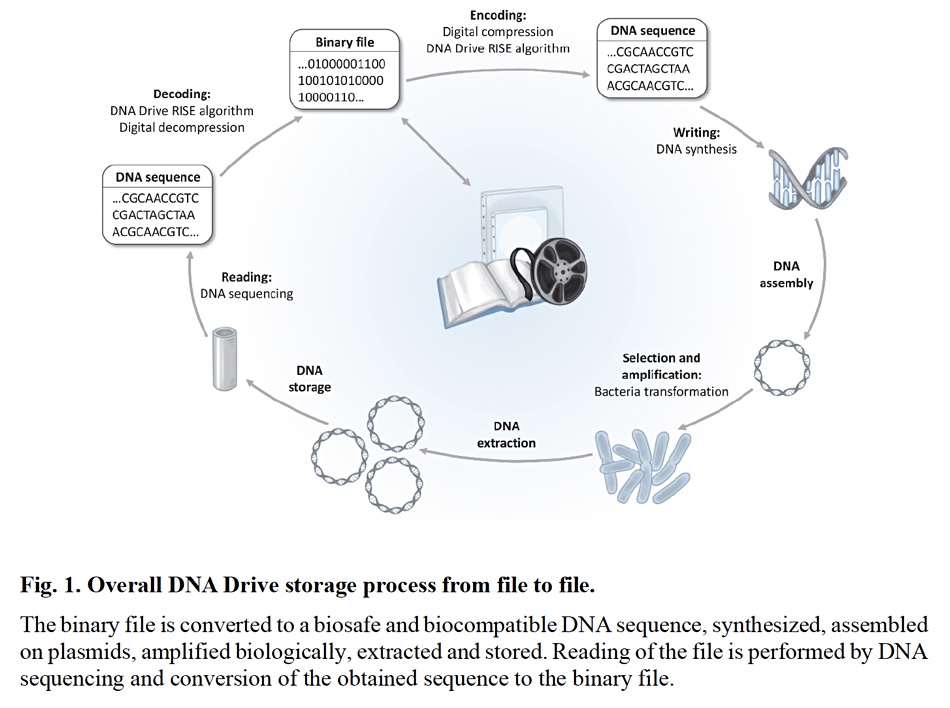
The read process necessitates locating the right capsule, unsealing it, rehydrating the DNA, extracting a droplet, putting it into an Oxford Nanopore MinION sequencer, retrieving the coded data, decompressing it and outputting the reconstructed digital file. Test capsules have been read multiple times with 100 percent fidelity.
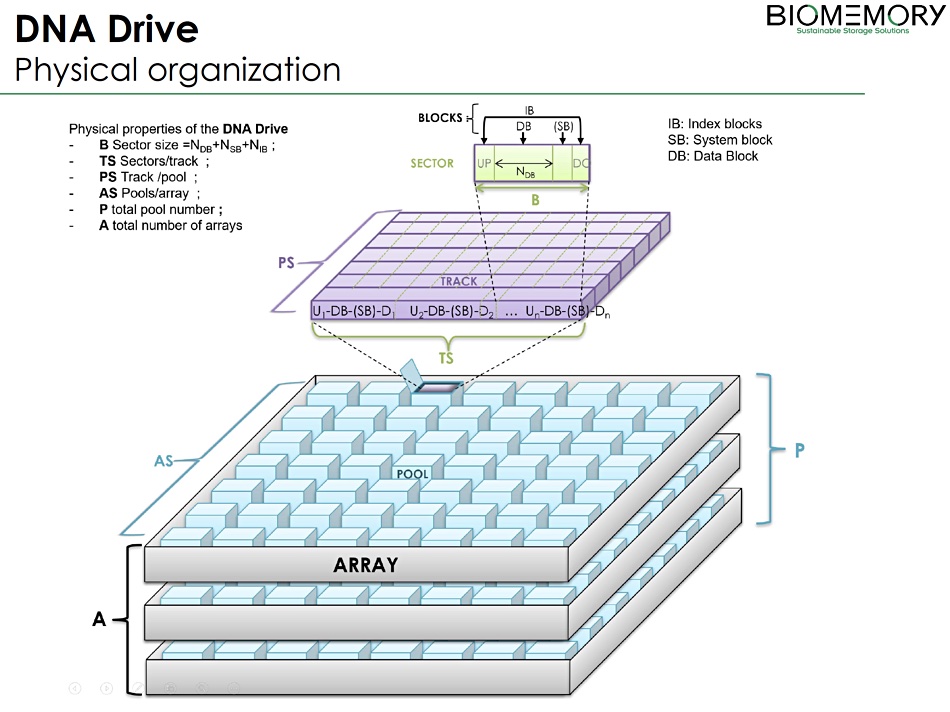
The DNA Drive has a file system ordering data in physical DNA sectors which supports metadata and partitions, allocation table, file/directory constructs and indexes. File data can be compressed, error code corrected and randomly accessed.
Biomemory has devised a DNA Drive concept organised as an array with pools of storage, organised into tracks (circular replicative molecules) and sectors, analogous to a disk drive, with index, system and data blocks in a track sector. The DNA Drive is sequenced at the pool level to recover data.
It has also envisioned a rackable DNA Data Storage Server 4RU high with removable DNA Drive cartridges and DNA “ink” cartridges, and a supposed $1/TB cost.
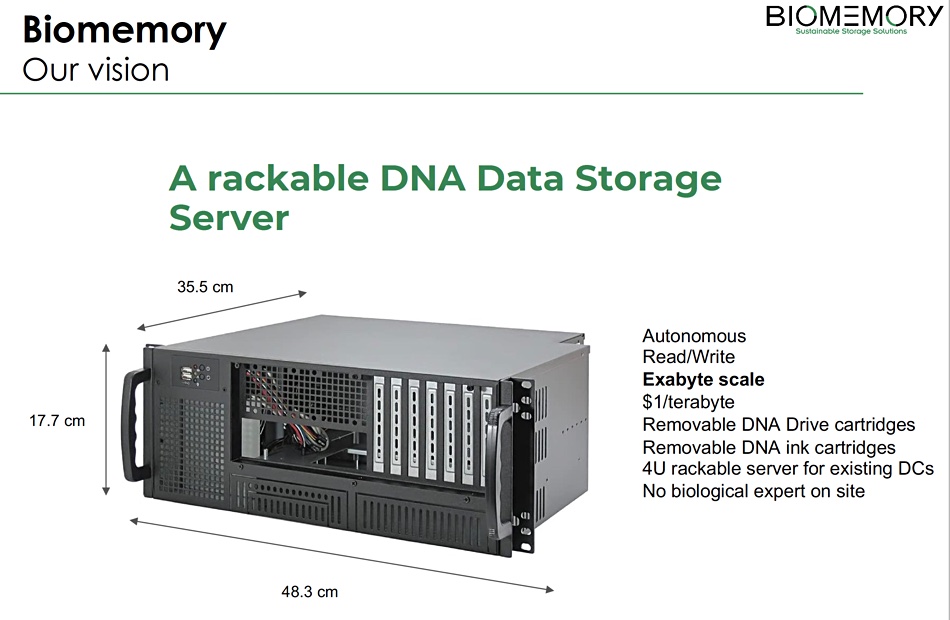

Why so big? Some number of 5PB pellets would fit into a cartridge and there would be some number of cartridges. Without knowing the numbers we don’t know this chassis’ capacity. There has to be a robot, a pretty fine-fingered robot, to move pellets between cartridges and an on-board sequencer. There has to be an embedded server to create, contain and manage the metadata indexing the pellets and their contents, otherwise random access would be truly random – you wouldn’t know what you are reading.
The metadata could be huge. The data/metadata ratio could vary between 1,000:1 for large files and down to even 1:10 with very small files. At 1,000:1 we would need 5TB of metadata to handle 5PB of data. That means a 1,000-pellet Data Storage Server would need 5PB of storage just to handle the metadata. And that storage would need to be online so that the unit could respond to cartridge load, unload and data read requests. The pellet capacity in any chassis is going be limited by the need for on-board metadata storage and compute capability – unless that was provided by an external server.
Bio-safety
The synthesized DNA is bio-compatible, in that it can be inserted into e coli and replicated as it reproduces, but also bio-safe. That means it has sections removed that could actually be used by the e.coli cells as they reproduce and so alter their genetic makeup. It is also encrypted so that it does not carry visible generic information.
Biomemory says bio-compatibility and bio-safety are ensured through its RISE (Random Iterative In-Silico Evolution) algorithm.
Background
Biomemory was founded as a spin-off from the Sorbonne Université and the French National Centre for Scientific Research (CNRS) in July 2021 by two academic researchers, CTO Pierre Crozet and CSO Stéphane Lemaire, and an entrepreneur, CEO Erfane Arwani.

The company is supported and funded by the Sorbonne University, the CNRS and technology development accelerator Satt Lutech, and also by external investors, including data storage businesses. Its technology partners include Twist Bioscience in San Francisco for DNA synthesis and Imagene in Bordeaux, France, for the DNA encapsulation in metal pellets (DNAShell minicapsules).
Biomemory says DNA Drive technology should have a negligible carbon footprint. It has two patents in France: EP193062478 and PCT EP2020/077497.
Read more about the DNA Drive technology in a translated scientific paper La révolution de l’ADN: biocompatible and biosafe DNA data storage.
Other startups and organisations developing DNA storage include Catalog, Georgia Tech Research Institute, Northwestern University, China’s Tianjin University,
A DNA Storage Alliance has been set up to develop a DNA Storage ecosystem. Members include Western Digital, Microsoft, Twist Bioscience and Illumina and others, including Biomemory.








