Netflix has engineered a topical tech twist to rev up interest in the Biohackers TV series – using DNA storage, claimed to last for a thousand years or more, to hold a record of the episodes.
Netflix asked ETH Zurich Professor Dr Robert Grass for help in getting the job done. He worked with Twist BioScience, a San Francisco-based synthetic biology company, which supplied the physical synthetic DNA segments.
Emily Leproust, CEO of Twist, issued a quote: “It’s exciting to ground the fictional series, which expounds beyond the boundaries of what is possible with DNA today, with the reality of preserving groundbreaking cultural media in synthetic DNA. The ability to store digital data in DNA seems futuristic, but the future is now.”
Grass said: “Theoretically in a gram of DNA you can pack 200 exabytes of data – that is 200 million terabytes. Theoretically, 20 grams of DNA would be enough to store all of the world’s digital data.”
A Youtube video explains the process.

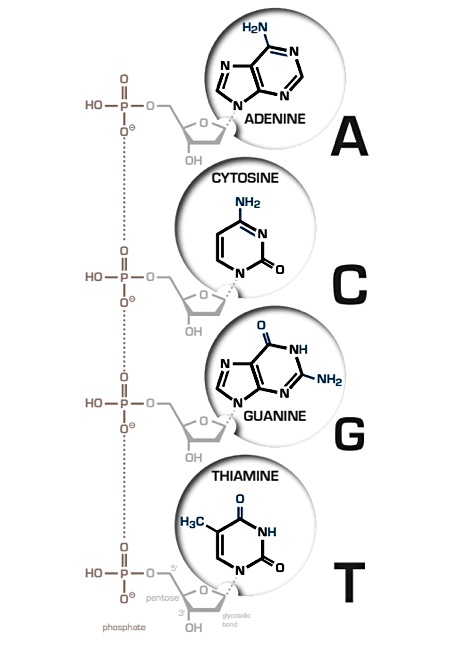
The Biohackers video series digital file is encoded into synthetic DNA. This comprises a DNA sequence of nucleic bases – adenine (A), guanine (G), cytosine (C) and thymine (T) – along with error-correction codes. For example, 00 = A, 01 = C, 10 = G and 11 = T. Each base can encode two bits of digital data. Sequential parts of an episode are encoded this way and index data is added to identify the location of each part in the episode.
The resulting data is encoded into short segments of DNA, 200 to 300 bases long, which are packed onto small, nanometre-sized, silica glass beads for preservation. Approximately four million segments were needed. The beads are put into a pink liquid, the liquid into a glass phial, and the phial stored. That’s the writing process.

Why use glass beads? It is because they resemble synthetic fossils, Grass said. “Hard drives and SD cards … are only stable for five to 20 years. But we have ancient fossil records, which are thousands of years old, and they still contain the natural fossil DNA.”
To read the data the DNA segments are washed off the glass beads and sequenced to reveal the string of A, G, C, and T sequences in each segment. The code strings are reconstituted into a digital file which can play the Biohackers episode. The entire read process takes about 24 hours.
A Twist Bioscience spokesperson said: ”There are a variety of ways to code/decode the data, so an end user can independently decide what method to use or ask a service provider like Twist to use their standard algorithms. Synthesis and sequencing are standardised, so, once again, an end user can decide who synthesises and sequences the DNA or they can subscribe to an end-to-end service, which Twist currently provides.”
She said: “There are also options for storing the synthesized DNA: it can be attached to glass beads and encased in silica; stored freeze-dried in a glass-lined, hermetically sealed tiny metal cylinder; or added to water, placed in a plastic tube and put into a -80C freezer.
“The retrieval process, required before sequencing, is easy when it is stored in a freezer, a bit more complicated when retrieved from a metal canister, and more complicated when retrieved from glass beads. The trade-off for the three storage systems is density/compactness versus retrieval complexity, with glass bead storage being the densest and most complicated retrieval.”
Twist says it can store one million small pieces of DNA on a silicon chip and is working towards upping that to 10GB of DNA on a chip. A Twist white paper says: “We see the potential for DNAbased storage to exceed bandwidths for writing disk and tape, but with dramatically smaller physical space and energy requirements and greatly increased stability.”
Also: “We anticipate that DNA storage systems in the short term will exceed the performance and cost characteristics of tape while retaining the advantages specific to DNA-based technology. From there, we expect another generation of technology to be mature enough to approach hard drive characteristics while maintaining the previous trajectory in cost advantage. “








